AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
Back to Blog
Peptide backbone of a protein8/8/2023  
geometry of the polypeptide backbone and on its experimental. Moreover, values for the "blocking effect" of side chains on the hydrogen exchange rates of peptide NH protons are correlated with ESF values. study confirms the deviation of s angles in both the peptide and protein structures but. As we learned in the Chapter 2, hydrogen bonds are directional (exhibiting a bond angle of 180) therefore, the peptide backbone must adopt specific conformations in order for atoms in peptide bonds to hydrogen bond with each other. The coil library values of $\langle \rangle $, which vary strikingly among the amino acids, are correlated with the neighboring residue effect seen by ESF. When tripeptides from the coil library are excised to allow solvation, the central residues have more negative values of 〈φ〉 but less negative values of 〈ESF〉 with L than with S neighbors. Several citrullinated proteins are associated with RA, suggesting that ACPAs may react with different sequence patterns, separating them from traditional. We show that L amino acids reduce the solvation of neighboring peptide groups more than S amino acids. The next level of protein structure, secondary structure, refers to local folded structures that form within a polypeptide due to interactions between atoms of the backbone. Calculated values of peptide solvation (electrostatic solvation free energy, ESF) predict basic properties of the neighboring residue effect. It is widely accepted that protein structures are made from low-energy parts (9), and consequently peptide. Detailed descriptions of GaMD theory and energetic reweighting were shown. When a neighboring residue (i - 1 or i 1) belongs to class L (aromatic and β-branched amino acids, FHITVWY) rather than class S (all others, G and P excluded), then the backbone angle φ of residue i is more negative for essentially all amino acids. The RMSDs of the peptide and protein backbone were used as reaction coordinates. Purchase Synthetic and Enzymatic Modifications of the Peptide Backbone, Volume 656 - 1st Edition. The neighboring residue effect appears also in the "coil library" of Protein Data Bank structures of residues not in α-helix and not in β-structure. This method would allow a breakthrough for the discovery of protein mimetic for. There is a neighboring residue effect on backbone conformation, discovered in 1997 from measurements of 3 J HNα coupling constants. Dynamic Amino Acid Side-Chains Grafting on Folded Peptide Backbone. 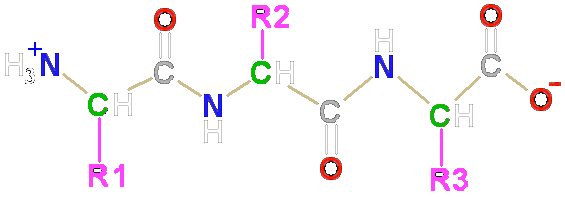
Unfolded peptides in water have some residual structure that may be important in the folding process, and the nature of the residual structure is currently of much interest. Protein structure depends on its amino acid sequence and local, low-energy chemical bonds between atoms in both the polypeptide backbone and in amino acid side chains. 
0 Comments
Read More
Leave a Reply. |
 RSS Feed
RSS Feed
